 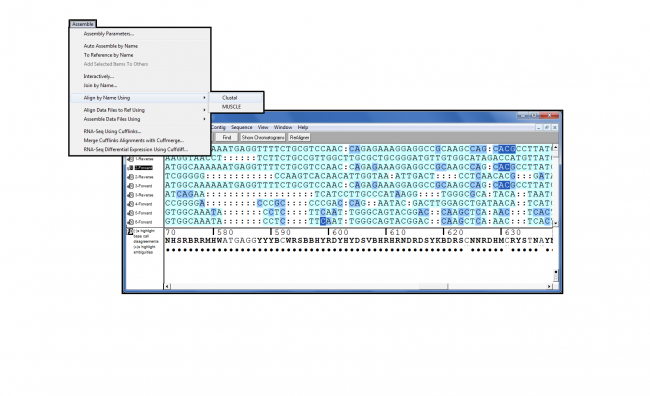
Acc Len: the number of nucleotides or amino acids in the result sequence identified by the accession number.Ident: the highest percent identity for a set of aligned segments to the same subject sequence.The expect value is the default sorting metric for significant alignments the E value should be very close to zero. E Value: the number of alignments expected by chance with the calculated score or better.Query Cover: the percent of the query length that is included in the aligned segments.Total Score: the sum of alignment scores of all segments from the same subject sequence.Max Score: the highest alignment score calculated from the sum of the rewards for matched nucleotides and penalities for mismatches and gaps.Note that the first match is a synthetic construct (that is, the sequence was computationally derived and is not associated with any organism): View the Descriptions tab to see a list of significant alignments. Under the Alignments tab next to Alignment view select Pairwise with dots for identities. This is an unknown protein sequence that we are seeking to identify by comparing it to known protein sequences, and so Protein BLAST should be selected from the BLAST menu:Įnter the query sequence in the search box, provide a job title, choose a database to query, and click BLAST: To access BLAST, go to Resources > Sequence Analysis > BLAST: Protein and gene sequence comparisons are done with BLAST (Basic Local Alignment Search Tool). There are also standalone and API BLAST options as well as pre-populated specialized searches available on the BLAST homepage linked above. This is useful when trying to identify a protein (see From sequence to protein and gene below). BLASTp (Protein BLAST): compares one or more protein query sequences to a subject protein sequence or a database of protein sequences.The HTG sequences, draft sequences from various genome projects or large genomic clones, are another large source of unannotated coding regions. Hence a tblastn search is the only way to search for these potential coding regions at the protein level. Since ESTs have no annotated coding sequences, there are no corresponding protein translations in the BLAST protein databases. They comprise the largest pool of sequence data for many organisms and contain portions of transcripts from many uncharacterized genes. ESTs are short, single-read cDNA sequences. Tblastn is useful for finding homologous protein coding regions in unannotated nucleotide sequences such as expressed sequence tags (ESTs) and draft genome records (HTG), located in the BLAST databases est and htgs, respectively. tBLASTn (protein sequence searched against translated nucleotide sequences): compares a protein query sequence against the six-frame translations of a database of nucleotide sequences.Thus blastx is often the first analysis performed with a newly determined nucleotide sequence. Because blastx translates the query sequence in all six reading frames and provides combined significance statistics for hits to different frames, it is particularly useful when the reading frame of the query sequence is unknown or it contains errors that may lead to frame shifts or other coding errors. 
BLASTx (translated nucleotide sequence searched against protein sequences): compares a nucleotide query sequence that is translated in six reading frames (resulting in six protein sequences) against a database of protein sequences.This is useful when trying to determine the evolutionary relationships among different organisms (see Comparing two or more sequences below). 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
